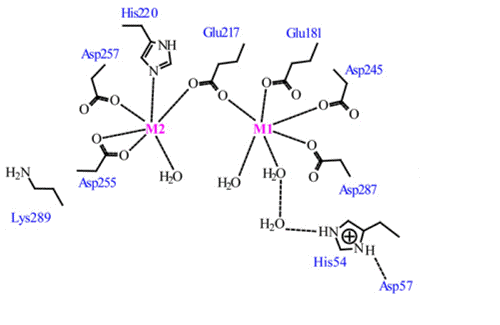
L-Arabinose Binding, Isomerization, and Epimerization by D-Xylose Isomerase: X-Ray/Neutron Crystallographic and Molecular Simulation Study: Structure

Locating active-site hydrogen atoms in d-xylose isomerase: Time-of-flight neutron diffraction | PNAS
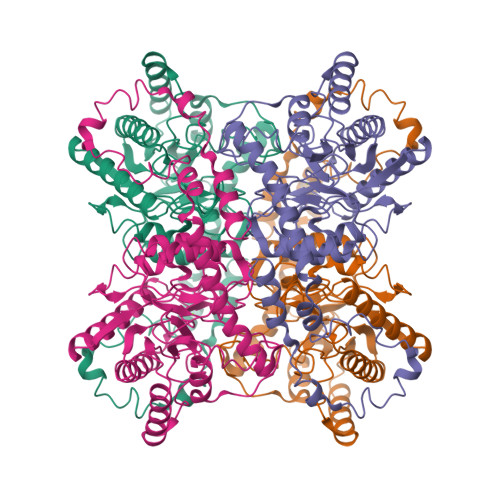
RCSB PDB - 2XIS: A METAL-MEDIATED HYDRIDE SHIFT MECHANISM FOR XYLOSE ISOMERASE BASED ON THE 1.6 ANGSTROMS STREPTOMYCES RUBIGINOSUS STRUCTURES WITH XYLITOL AND D-XYLOSE

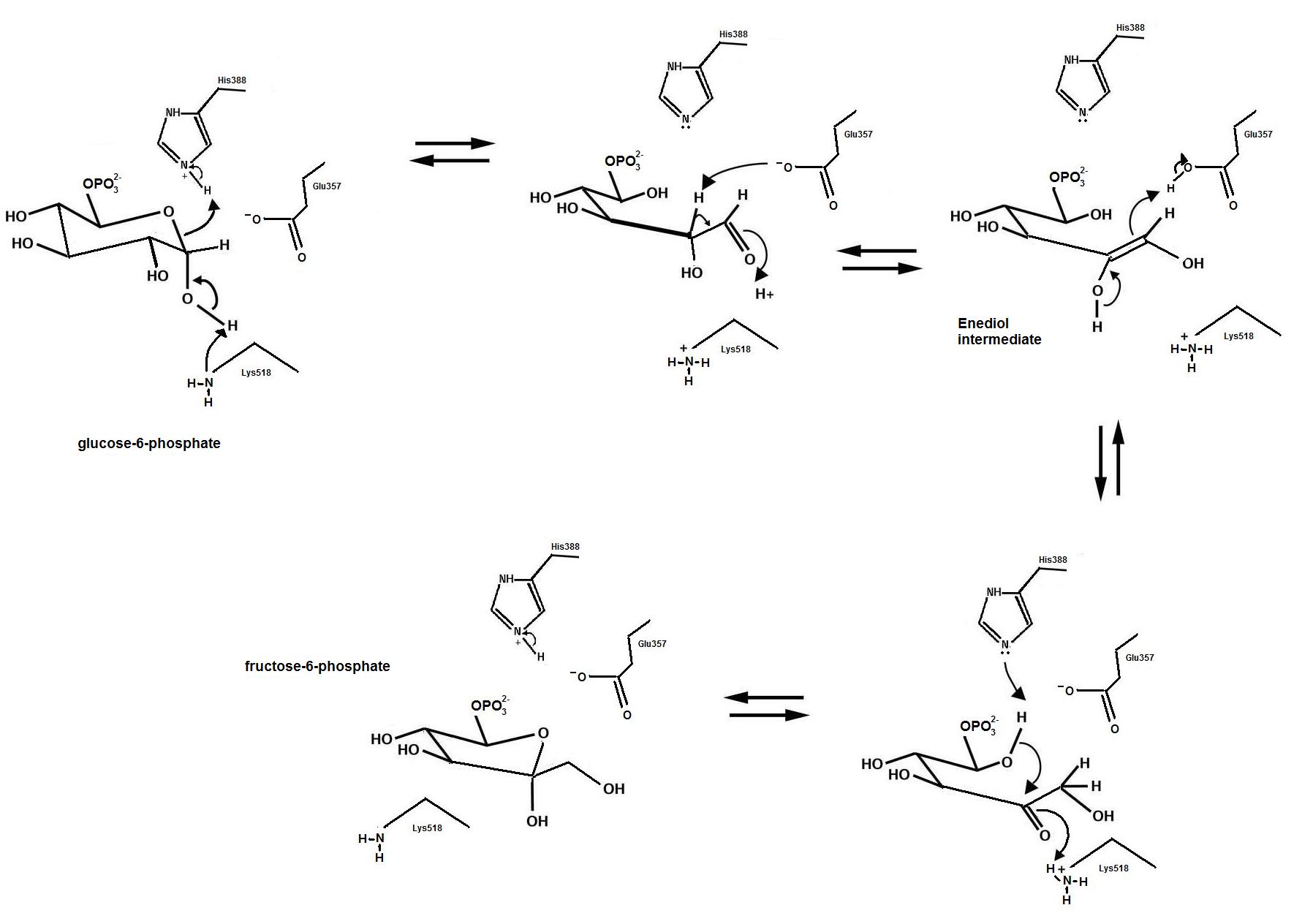
Catalytic mechanism of xylose isomerase. Top: ring opening, middle:... | Download Scientific Diagram

Locating active-site hydrogen atoms in d-xylose isomerase: Time-of-flight neutron diffraction | PNAS
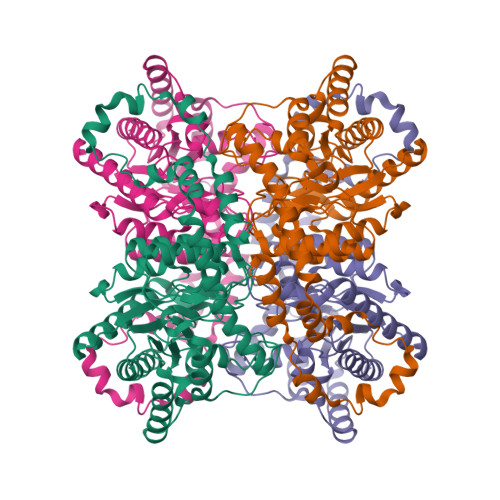

RCSB PDB - 8XIA: X-RAY ANALYSIS OF D-XYLOSE ISOMERASE AT 1.9 ANGSTROMS: NATIVE ENZYME IN COMPLEX WITH SUBSTRATE AND WITH A MECHANISM-DESIGNED INACTIVATOR

Xylose fermentation efficiency of industrial Saccharomyces cerevisiae yeast with separate or combined xylose reductase/xylitol dehydrogenase and xylose isomerase pathways | Biotechnology for Biofuels and Bioproducts | Full Text
Proposed catalytic reaction mechanism including sugar-ring openings of... | Download Scientific Diagram

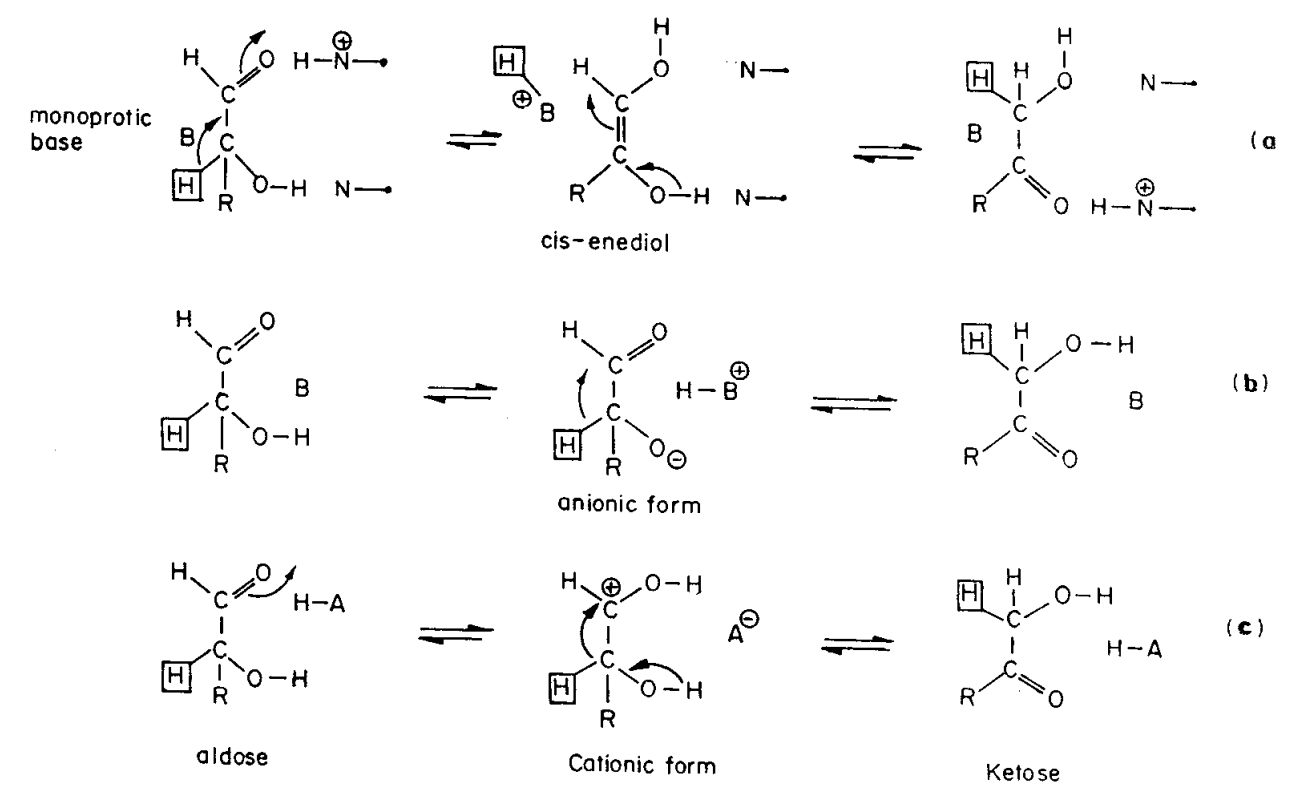
Figure 1 from Observations of reaction intermediates and the mechanism of aldose-ketose interconversion by D-xylose isomerase. | Semantic Scholar
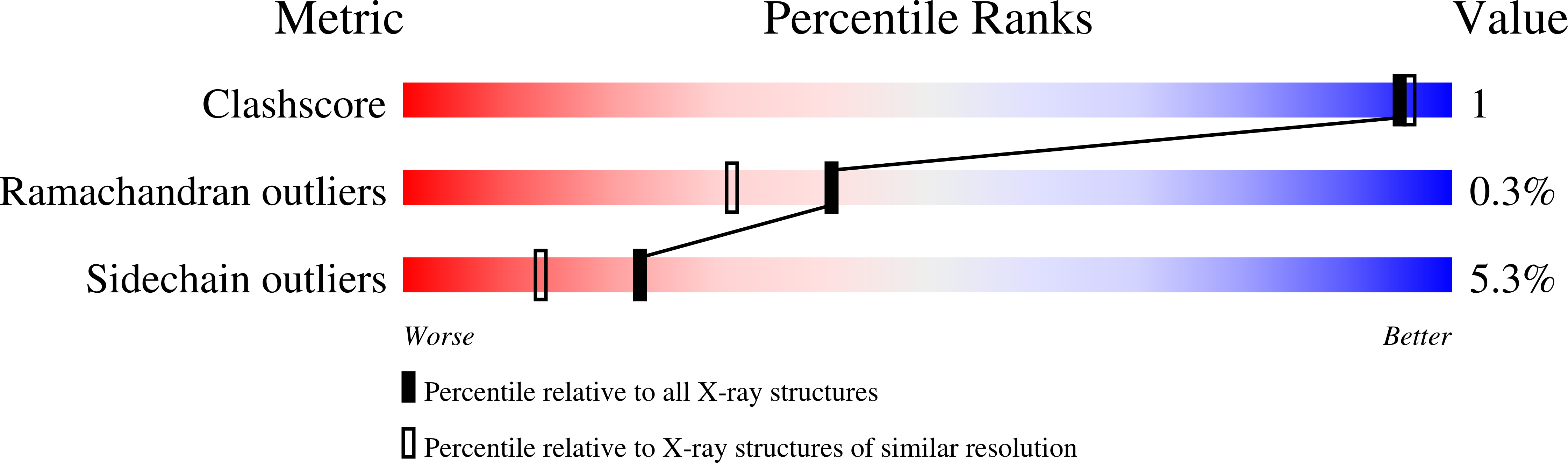
RCSB PDB - 8XIA: X-RAY ANALYSIS OF D-XYLOSE ISOMERASE AT 1.9 ANGSTROMS: NATIVE ENZYME IN COMPLEX WITH SUBSTRATE AND WITH A MECHANISM-DESIGNED INACTIVATOR

L-Arabinose Binding, Isomerization, and Epimerization by D-Xylose Isomerase: X-Ray/Neutron Crystallographic and Molecular Simulation Study: Structure
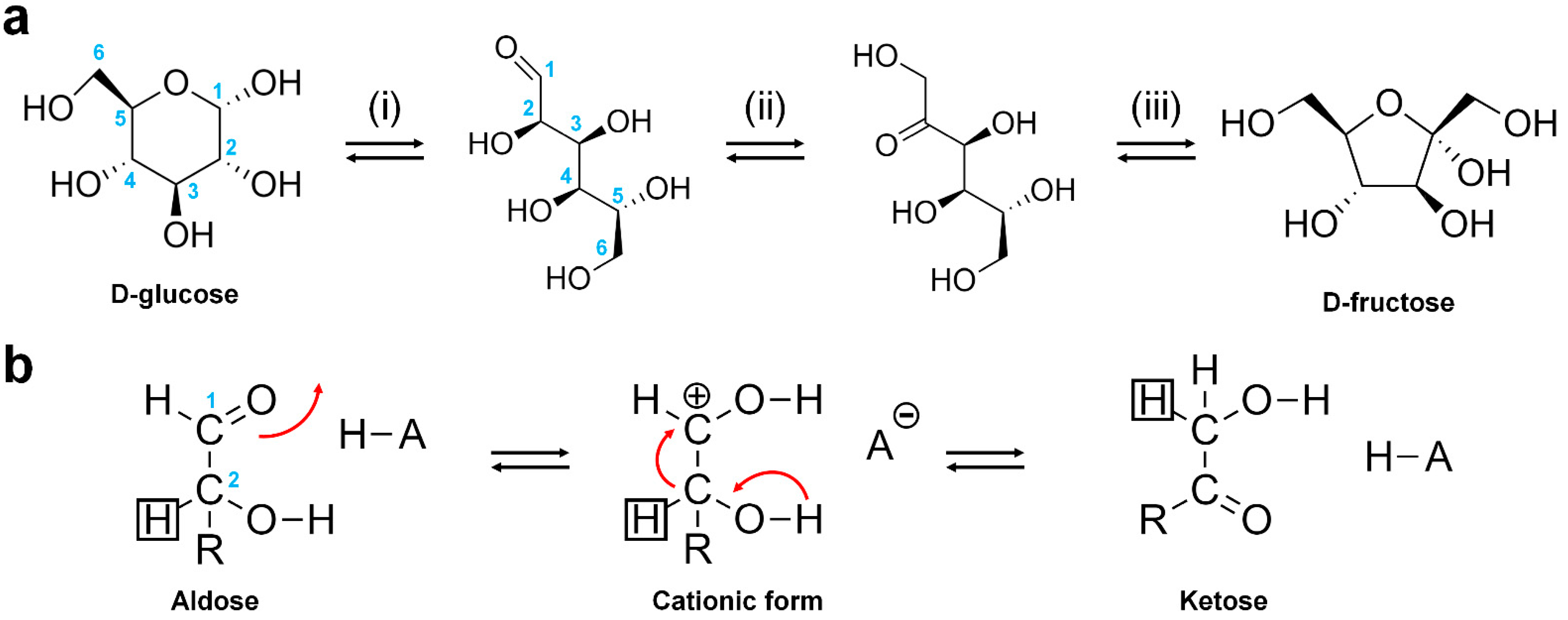
Applied Sciences | Free Full-Text | Glucose Isomerase: Functions, Structures, and Applications | HTML